0. Current research: My current work leverages deep learning tools (e.g. simple MLPs, CNNs, and large language models), to answer questions from genomic data that cannot be easily directly inferred from simpler models. This is particularly handy in population genetics where extremely efficient and robust simulators like msprime allow us to produce synthetic training data for our models. I'm currently using these approaches to flexibly infer certain population genetic parameters from whole genomes, and exploring their use in understanding antibiotic resistance.
|
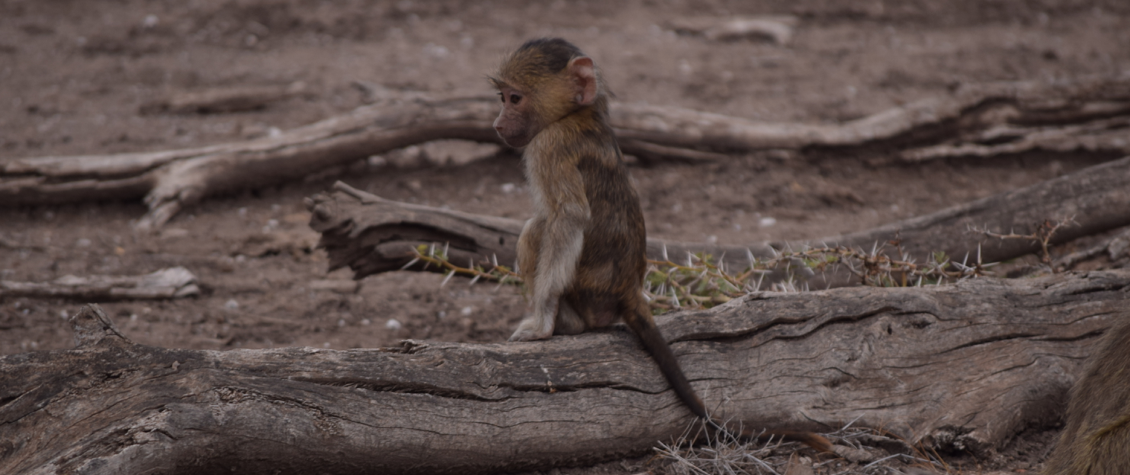
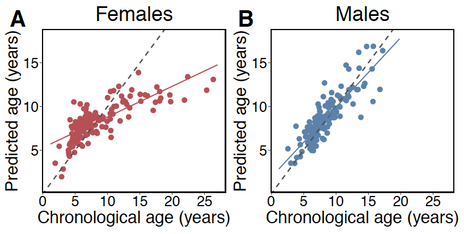
1. Epigenetic clocks: DNA methylation underlies many aspects of aging. Increasingly genome-wide DNA methylation patterns are being used to estimate chronological ages, deviations from which may provide insight into inter-individual differences in biological aging. These properties of "epigenetic clocks" can be used to ask about socio-environmental factors that shape aging. We applied these methods for the first time in a wild primate population to show that high ranking male baboons show accelerated epigenetic ages (Anderson et al. 2021, eLife). We are continuing work to see whether these results extend to other species, including those with different rank dynamics.
|
|
2. Early life adversity and DNA methylation: The early life environment experienced by an animal (both social and non-social) has large and often long-lasting effects on lifespan. However we understand little about the molecular mechanisms that mediate this relationship. This project aims to address this gap by exploring the relationship between adversity in early life and adulthood, and adult DNA methylation patterns in the Amboseli baboons. See our new preprint.
|
|
3. Social environment and the immune response: Work in captive primates where social status can be manipulated has demonstrated a clear, strong causal relationship between social status and gene regulation within the immune system (Snyder-Mackler et al. 2016, Science). This project extends this, and other work, to explore how social status and social integration combine to shape immune gene regulation, including in response to mimicked pathogen threat. This work is now available as a preprint (doi: https://doi.org/10.1101/2021.05.31.446340).
|
|
4. Applications for field-based sequencing technology: Portable sequencing technology, like that developed by Oxford Nanopore, has a wide-range of attractive applications in long-term field studies. However, because of the recency of their development, and the pace of their advancement, many of these applications have yet to be realized on a wide scale. This work explores one such application, which is a method for cheap, low-resource parentage assignment from non-invasively collected samples in the field.
|